Sequencing Parulidae, a Rapid Avian Radiation
In 2016, we sequenced the first high quality reference genome of a myrtle warbler. In 2021, we updated this with a chromosome-level re-assembly. This has supported numerous projects, both for our own research as well as the broader avian genomics community. We are now interested in mapping the genetic basis of divergent phenotypes across many species in the family and quantifying evolutionary patterns amongst this group of birds! Affectionately known as the “Parulome Project”, we aim to sequence multiple individuals from every species in the radiation (>100). This work is currently supported by a NSF grant to PI Toews.
Genomic Divergence and the Consequences of Hybridization
Along with colleagues at the Cornell Lab of Ornithology, the University of Colorado Boulder, the University of British Columbia, and UC—Riverside, we are applying genomic tools (i.e. whole-genome re-sequencing and reduced representation sequencing) to the golden-winged / blue-winged warbler system and the yellow-rumped warbler system. We are interested in learning about the dynamics of hybridization, the patterns of genomic divergence, and understanding the genetic basis for phenotypic differences. Hybridization and gene flow likely play a key role in the evolutionary histories of these and other species groups, and we are keen to understand the kind and extent of gene sharing between divergent lineages.
Genetics of Coloration
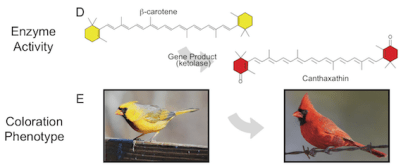
Empowered by findings of candidate feather coloration and patterning genes in warblers, we are beginning to understand the important genetic players that generate the amazing phenotypic diversity of birds. Our current focus is to better understand the genes involved in carotenoid processing. In most animals, carotenoids must be acquired from the environment and are not water-soluble and therefore require special transport to tissues for deposition. Unlike genes involved in melanin production, we know much less about the genetic basis of carotenoid coloration, and our research is trying to highlight them.
Parulid warblers are not only interesting from an evolutionary genomics perspective. Since Robert MacArthur’s early work on the group in the 1960s, the species have also raised questions about niche divergence and competitive exclusion in the world of community ecology. We have approached these questions using molecular data, which include fecal metabarcoding to learn about diet, microbiome, and have collaborated on work studying the evolution of their feather mites. Some of this work is currently supported by a NSF CAREER grant to PI Toews.
Mitochondrial Introgression
Different genetic markers can tell us different evolutionary stories. I have been particularly interested in the drivers of discordant patterns between mitochondrial and nuclear DNA. Empirically I have studied this in Audubon’s warblers in the Southwestern U.S. This is an area where previous studies suggested there is a cryptic contact zone between divergent mitochondrial clades, one introgressed from a close relative—myrtle warblers—and the other the assumed ancestral type that occurs primarily in black-fronted warblers. Along with collaborators I used biogeographic patterns, genotyping, stable isotopes and mitochondrial biochemistry to characterize the birds in this region to better understand the drivers of this instance mitochondrial introgression.
Migration in an Evolutionary Context
Many migrating animals travel great distances between their wintering and breeding grounds. In many species, the process of migrating between these distant areas is both a navigational and energetic challenge. How do they accomplish this feat and, given that past research suggests a strong genetic basis, how do differences evolve, what are the important genes, and how does hybridization contribute? To better understand this I have used behavioural trials (i.e. orientation experiments), stable isotope, biogeographic, and citizen science data to understand the migratory patterns of several taxa, including some that hybridize.
Hybridization in Suture Zones
Areas where numerous taxa come into contact at their range margins are known as “suture zones”. Studying the extent of hybridization and reproductive isolation in these areas can teach us a lot about the process of speciation. We have studied one such area in northeastern B.C., along the eastern foothills of the Rocky Mountains. In several species we have quantified possible admixture in both genetic, behavioural (e.g. song), and other phenotypic traits.
Reproductive Isolation and Cryptic Diversity
Cryptic diversity is the kind of biological variation that can be difficult to identify, at least based on superficial phenotypic characters. For example, our study of a contact zone between divergent forms of winter wren—now split into two distinct species—show that plumage similarities belied strong differences in their genes and vocalizations. We used these kinds of data in an area in northeast B.C. to show that individuals of the two groups do not interbreed where they co-occur and, in combination with the large genetic differences noted between them, are likely reproductively isolated. These data suggest that there are possibly other species with deep genetic differences in the absence of strong phenotypic differences, and this has important implications for our ability to understand and quantify biodiversity.






